Demo for the DoWhy causal API
We show a simple example of adding a causal extension to any dataframe.
[1]:
import dowhy.datasets
import dowhy.api
import numpy as np
import pandas as pd
from statsmodels.api import OLS
[2]:
data = dowhy.datasets.linear_dataset(beta=5,
num_common_causes=1,
num_instruments = 0,
num_samples=1000,
treatment_is_binary=True)
df = data['df']
df['y'] = df['y'] + np.random.normal(size=len(df)) # Adding noise to data. Without noise, the variance in Y|X, Z is zero, and mcmc fails.
#data['dot_graph'] = 'digraph { v ->y;X0-> v;X0-> y;}'
treatment= data["treatment_name"][0]
outcome = data["outcome_name"][0]
common_cause = data["common_causes_names"][0]
df
[2]:
| W0 | v0 | y | |
|---|---|---|---|
| 0 | 0.076236 | True | 5.553473 |
| 1 | 0.060844 | False | 0.368952 |
| 2 | -0.028633 | False | -0.494726 |
| 3 | -1.427719 | True | 5.404953 |
| 4 | -0.671959 | False | -1.001133 |
| ... | ... | ... | ... |
| 995 | 0.700718 | True | 5.251570 |
| 996 | 0.946380 | True | 7.180262 |
| 997 | -0.004310 | True | 4.482246 |
| 998 | 0.618509 | True | 4.455502 |
| 999 | -1.381954 | False | -0.086505 |
1000 rows × 3 columns
[3]:
# data['df'] is just a regular pandas.DataFrame
df.causal.do(x=treatment,
variable_types={treatment: 'b', outcome: 'c', common_cause: 'c'},
outcome=outcome,
common_causes=[common_cause],
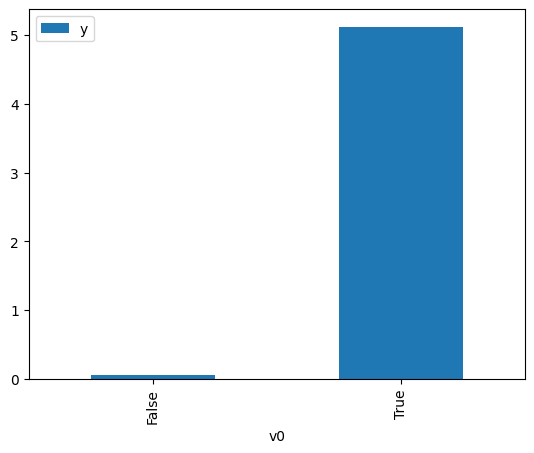
proceed_when_unidentifiable=True).groupby(treatment).mean().plot(y=outcome, kind='bar')
[3]:
<AxesSubplot: xlabel='v0'>
[4]:
df.causal.do(x={treatment: 1},
variable_types={treatment:'b', outcome: 'c', common_cause: 'c'},
outcome=outcome,
method='weighting',
common_causes=[common_cause],
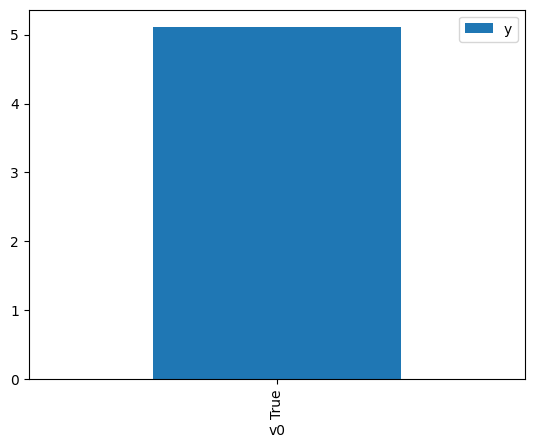
proceed_when_unidentifiable=True).groupby(treatment).mean().plot(y=outcome, kind='bar')
[4]:
<AxesSubplot: xlabel='v0'>
[5]:
cdf_1 = df.causal.do(x={treatment: 1},
variable_types={treatment: 'b', outcome: 'c', common_cause: 'c'},
outcome=outcome,
dot_graph=data['dot_graph'],
proceed_when_unidentifiable=True)
cdf_0 = df.causal.do(x={treatment: 0},
variable_types={treatment: 'b', outcome: 'c', common_cause: 'c'},
outcome=outcome,
dot_graph=data['dot_graph'],
proceed_when_unidentifiable=True)
[6]:
cdf_0
[6]:
| W0 | v0 | y | propensity_score | weight | |
|---|---|---|---|---|---|
| 0 | -0.352919 | False | 0.446923 | 0.584806 | 1.709968 |
| 1 | 0.814069 | False | 0.333252 | 0.282563 | 3.539036 |
| 2 | -2.343394 | False | 1.050524 | 0.925262 | 1.080776 |
| 3 | 0.828998 | False | -0.382004 | 0.279270 | 3.580766 |
| 4 | 1.064405 | False | 0.701340 | 0.230561 | 4.337242 |
| ... | ... | ... | ... | ... | ... |
| 995 | 0.895503 | False | 0.163814 | 0.264889 | 3.775160 |
| 996 | 2.014276 | False | -0.379485 | 0.096009 | 10.415706 |
| 997 | -0.376085 | False | -0.769199 | 0.590935 | 1.692233 |
| 998 | 1.836714 | False | 1.371098 | 0.114205 | 8.756149 |
| 999 | 0.428957 | False | -0.634386 | 0.374900 | 2.667378 |
1000 rows × 5 columns
[7]:
cdf_1
[7]:
| W0 | v0 | y | propensity_score | weight | |
|---|---|---|---|---|---|
| 0 | 1.303760 | True | 5.675314 | 0.812527 | 1.230728 |
| 1 | 1.436695 | True | 6.111159 | 0.833644 | 1.199553 |
| 2 | 0.631619 | True | 6.052427 | 0.675366 | 1.480680 |
| 3 | 0.354007 | True | 6.152660 | 0.605733 | 1.650893 |
| 4 | -2.741986 | True | 2.913577 | 0.049674 | 20.131449 |
| ... | ... | ... | ... | ... | ... |
| 995 | 0.721921 | True | 5.594780 | 0.696601 | 1.435543 |
| 996 | 1.337709 | True | 3.280717 | 0.818109 | 1.222331 |
| 997 | -2.741986 | True | 2.913577 | 0.049674 | 20.131449 |
| 998 | 1.812561 | True | 5.540571 | 0.883099 | 1.132376 |
| 999 | 0.124867 | True | 5.605328 | 0.544679 | 1.835943 |
1000 rows × 5 columns
Comparing the estimate to Linear Regression
First, estimating the effect using the causal data frame, and the 95% confidence interval.
[8]:
(cdf_1['y'] - cdf_0['y']).mean()
[8]:
$\displaystyle 5.06489946748145$
[9]:
1.96*(cdf_1['y'] - cdf_0['y']).std() / np.sqrt(len(df))
[9]:
$\displaystyle 0.0887509312439617$
Comparing to the estimate from OLS.
[10]:
model = OLS(np.asarray(df[outcome]), np.asarray(df[[common_cause, treatment]], dtype=np.float64))
result = model.fit()
result.summary()
[10]:
| Dep. Variable: | y | R-squared (uncentered): | 0.942 |
|---|---|---|---|
| Model: | OLS | Adj. R-squared (uncentered): | 0.941 |
| Method: | Least Squares | F-statistic: | 8038. |
| Date: | Fri, 16 Dec 2022 | Prob (F-statistic): | 0.00 |
| Time: | 19:37:21 | Log-Likelihood: | -1410.4 |
| No. Observations: | 1000 | AIC: | 2825. |
| Df Residuals: | 998 | BIC: | 2835. |
| Df Model: | 2 | ||
| Covariance Type: | nonrobust |
| coef | std err | t | P>|t| | [0.025 | 0.975] | |
|---|---|---|---|---|---|---|
| x1 | 0.2405 | 0.036 | 6.761 | 0.000 | 0.171 | 0.310 |
| x2 | 5.0122 | 0.048 | 104.315 | 0.000 | 4.918 | 5.106 |
| Omnibus: | 1.561 | Durbin-Watson: | 1.932 |
|---|---|---|---|
| Prob(Omnibus): | 0.458 | Jarque-Bera (JB): | 1.428 |
| Skew: | -0.062 | Prob(JB): | 0.490 |
| Kurtosis: | 3.137 | Cond. No. | 1.94 |
Notes:
[1] R² is computed without centering (uncentered) since the model does not contain a constant.
[2] Standard Errors assume that the covariance matrix of the errors is correctly specified.